FASTQ
August 28, 2020
File Names
SampleName_S1_L001_R1_001.fastq.gz
- SampleName > sample name provided
- S1 > sample number based on order in list of samples. S1 is first sample on the list.
- L001 > Lane number
- R1 > Read number. Paired end sequencing must have at least one R1 file and 1 R2 file.
- 001 > The last segment is always 001.
Illumina FASTQ naming convention
Format
For a single-read run, one Read 1 (R1) FASTQ file is created for each sample per flow cell lane. For each cluster that passes filter, a single sequence is written to that samples’s R1 FASTQ file. So you can think of a single read as the sequence produced by a single cluster in one flowcell during one sequencing run.
Each read has 4 lines:
@EAS139:136:FC706VJ:2:2104:15343:197393 1:N:18:1
GATTTGGGGTTCAAAGCAGTATCGATCAAATAGTAAATCCATTTGTTCAACTCACAGTTT
+
!''*((((***+))%%%++)(%%%%).1***-+*''))**55CCF>>>>>>CCCCCCC65
1st Line
@<instrument>:<run_number>:<flowcell ID>:<lane>:<title>:<x-pos>:<y-pos>:<UMI> <read>:<is_filtered>:<control_number>:<index>
2nd Line
The sequence.
3rd Line
“+”. The space this line adds helped make the FASTQ format popular.
4th Line
- Quality scores for the read.
- Uses ASCII symbols to more efficiently encode multi-digit quality scores. See Encoding Basics
- Quality (!) means 33, which in a 33 based quality score system means a phred score of 0.
- Quality scores are usually 33 based because the first written ASCII character codes for 33.
- Some older quality score system are 64 based.
-
Phred scores are like this:
- 0: 0% confidence
- 10: 90% confidence and 1/10 error rate
- 20: 99% confidence and 1/100 error rate
- 30: 99.9% confidence and 1/1000 error rate
- 40: 99.99% confidence and 1/10000 error rate
- If sequencing gives a base call a phred score of 10, then the FASTQ file will show the ASCII character for 43 (33 + 10) which is +. If a base call gets a phred score of 20 that would be represented by the ASCII character for 53 (33 + 20) which is 5.
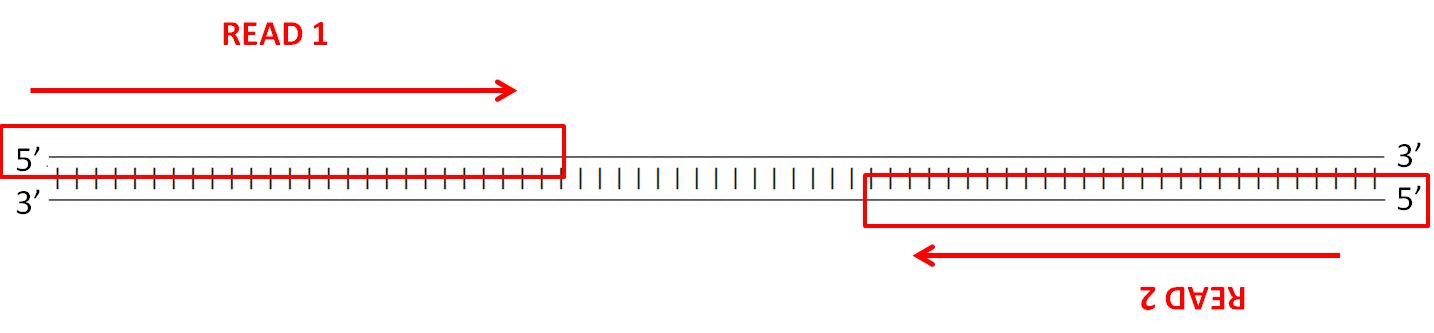
Forward and reverse reads